Drop Date: December 2015
SBB – Salivary DNA Genotyping
In This Drop: 1+1=3 (Biomarkers + DNA = Great Research)
The latest research continues to reinforce the correlation between SNP-related associations and genetic biology, behaviors, and diseases. But did you know that you can easily and affordably incorporate accurate, high-quality DNA analysis from saliva and get results that are identical to blood samples?
By optimizing the extraction process, Salimetrics created a protocol for researchers that optimizes high quality salivary DNA, using the same inexpensive saliva collection methods researchers have relied on for years. This enables researchers to expand the scope of their studies by including specific SNPs for genomic DNA testing without the need for specialized collection devices, preservatives or complicated collection techniques. For studies focused on a targeted number of SNPs, Salimetrics’ new protocol enables researchers to incorporate the efficiency and simplicity of using the same saliva sample for genetic analysis that researchers use for other biomarkers.
Empirical evidence shows that variations in gene composition (SNPs & VNTRs) express the ability to moderate behavioral tendencies, emotional resilience and responses, as well as vulnerability to disease states. SNP genotyping creates an inter-individual biological profile that enhances the conclusions of observed abnormalities in participant groups. For example, past studies on child maltreatment have shown a correlation between children with genetic variants in OXTR (specifically AA/AG) and being more at risk for increased emotional dysregulation when compared to children with the GG variant who were at lowest risk (Bakermans-Kranenburg et al., 2008; Rodrigues et al., 2009). When genomic analysis was used, there was a definite contrast to these past studies and it was discovered that maltreated children who carry AA/AG genetic variants were not more at risk for increased emotional dysregulation but were rather resilient individuals (Bradley 2011).
Bodenmann, 2009 studied subjects with the Val/Val genotype for the polymorphism of COMT and participants responded positively to Modafinil, a drug that stimulates wakefulness by modifying dopaminergic and nor-adrenergic neurotransmission. Results of this study confirmed that participants with the COMT polymorphism were influenced by Modafinil, which allowed them to function properly after a sleepless night and how the Met/Met genotype subjects were hardly affected by the drug. The use of DNA analysis this study became the first instance where a drug intervention yielded high and stable waking functions for a genetically distinct group with sleep deprivation.
Frielingsdorf et al., 2010 observed that carriers of the BDNF Met allele were capable of extinguishing a learned fear memory and fMRI proved these individuals were slower at suppressing a learned fear response. Frielingsdorf further discusses the importance of BDNF genotyping and neuroimaging when it comes to customizing a therapeutic treatment for PTSD.
Way et al., 2008 used genotyping in a study to provide a solid case for the association between OPRM1 and both physical and social pain analysis. In this study, DNA Analysis of the OPRM1 gene was used to show that the same neural and neurochemical systems behind generating the uncomfortable feelings of physical pain are also in charge of creating feelings of “social pain” (hurt feelings that result from social rejection, separation, or loss). Through fMRI it was shown in areas of the brain that have previously been proven to associate with social and physical pain, that G allele carriers had a greater reaction to social rejection.
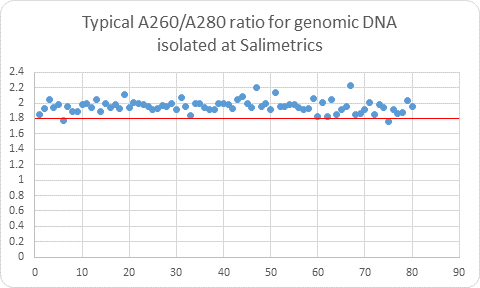
The SalivaLab’s DNA isolation protocol yields high purity, low salt DNA with A260/A280 ratios (indicating protein content) consistently at or above 1.8 and the A260/A230 ratios (indicative of salt and carbohydrate content) at or above 1.5. These high quality results with low sample retest rates allow for confidence in SNP calls and exceedingly low non-reportable findings (0.05%). The final result is a cost-effective service for targeted DNA genotyping, bringing ease, affordability, and painless sample collection to researchers. You already have access to the genetic material from your existing samples that you have collected for other analytes, and now with the simplicity of Salimetrics SalivaLab you have the opportunity to gain additional insight with cost-effective genetic analysis.
REFERENCES & RELATED RESEARCH
- Bakermans-Kranenburg, M., & van IJzendoorn, M (2008). Oxytocin receptor (OXTR) and serotonin transporter (5-HTT) genes associated with observed parenting. Social Cognitive and Affective Neuroscience, 3(2), 128–134.
- Rodrigues, S. M., Saslow, L. R., Garcia, N., John, O. P., & Keltner, D (2009). Oxytocin receptor genetic variation relates to empathy and stress reactivity in humans. Proceedings of the National Academy of Sciences of the United States of America, 106(50), 21437–21441.
- Bradley, B. et al (2011). Association between childhood maltreatment and adult emotional dysregulation in a low-income, urban, African American sample: Moderation by . Development and Psychopathology, 23(2011), 439-452.
- Bodenmann, S. et al (2009). Pharmacogenetics of Modafinil After Sleep Loss: Catechol-O-Methyltransferase Genotype Modulates Waking Functions But Not Recovery Sleep. Nature, 85(3), 296-304.
- Frielingsdorf, H. et al (2010). Variant brain-derived neurotrophic factor Val66Met endophenotypes: implications for posttraumatic stress disorder. Annals of the NY Academy of Sciences, 1208, 150-157.
*Note: Salimetrics provides this information for research use only (RUO). Information is not provided to promote off-label use of medical devices. Please consult the full-text article.
 Contact: Salimetrics (USA)
Contact: Salimetrics (USA)